Protecting crops from fungal diseases is often described as a never-ending arms race since pathogens continuously evolve to break down resistance genes. Now the drivers of that process are being used to simulate pathogen evolution in computer models that can predict the impacts of different disease control strategies.
The project was undertaken by Dr Peter Thrall and Dr Luke Barrett of CSIRO Agriculture and Food. The computer models were built specifically for disease control research in collaboration with the French National Research Institute for Agriculture (INRA) and postdoctoral researcher Dr Loup Rimbaud.
GRDC investment made it possible to set up this predictive technology, starting with Blackleg in canola and Stripe rust in bread wheat. In dollar terms, these are the two biggest threats to broadacre grains production in Australia.
 Stripe rust (Puccinia striiformis f. sp. tritici) affecting both leaf and heads in a susceptible variety in a stripe rust nursery. Photo: Luise Sigel
Stripe rust (Puccinia striiformis f. sp. tritici) affecting both leaf and heads in a susceptible variety in a stripe rust nursery. Photo: Luise Sigel
However, the model is broadly adaptable to other cropping systems.
Interpreting the outputs, however, is technical and requires statistical frameworks. That means the software is primarily a research and management tool and is not initially intended for on-farm use.
Included in the model is the ability to simulate crop growth and seasonal climatic conditions. It allows for spatial analysis within a two-dimensional landscape to alter variables such as pathogen dispersal.
“The model can account for variation in many pathogen traits,” Dr Barrett says. “Plus, it is stochastic, meaning we can simulate the natural variation present within pathogen populations. So even with the same set of input parameters, we never get exactly the same result, just like in the paddock.”
For the past three years it has been used to run thousands of simulations exploring the impacts of different strategies for deploying disease resistance genes.
This entailed using the model to look at how the pathogens behave under different approaches to crop resistance – the use of major and minor resistance genes, pyramiding of resistance genes, different spatial and temporal arrangements, crop rotations, or mixes of different strategies.
These were analysed in terms of how quickly the pathogen evolves to overcome resistances and also the severity of disease epidemics under different conditions.
The scale of analysis possible with modelling far exceeds the capacity of field trials to explore these issues. As such, this project allowed the first in-depth deconstruction of the complexity of the host-pathogen arms race against resistance genes.
“There are too many variables and combinations to study experimentally at landscape scales,” Dr Thrall says. “To find your way through all that complexity, you need modelling. That’s not to say you don’t need the real-world work; you absolutely do. Everything that comes out of the models has to be ground-tested. But the modelling can guide the empirical research.”
In fact, the project ran field trials for the past three of the project’s five years to test the model predictions in both canola and wheat. Dr Barrett also ran glasshouse and genetic tests to delineate important pathogen characteristics, such as aggressiveness, to ensure the model’s accuracy.
Challenging assumptions
Dr Barrett says the project tested all the big ideas relating to disease control using resistance genes.
Included is the assumption that gene pyramiding (also known as gene stacking) is the best strategy. This technique combines multiple disease resistance genes in the same plant to raise the hurdle the pathogen has to jump to break down that resistance.
“There was little systematic science underlying that assumption,” he says. “So that was one of the key questions we were looking at with the model. How do you use resistance gene diversity in a way that prolongs its effectiveness?”
The question has serious evolutionary implications. As with human pathogens that have acquired resistance to multiple antibiotics to create ‘superbugs’, evolutionary principles in the paddock raise the spectre of super-pathogens that can attack every variety.
“Part of our philosophy is to try and bring some of that evolutionary understanding into crop management and cropping landscapes,” Dr Thrall says.
When it comes to resistance genes, among the key findings are several important realisations:
- pyramids are not a universal answer to the host-pathogen arms race. They are good under some conditions, but not others. They are not a panacea;
- the assumption that minor resistance genes can protect major resistance genes is also not universally true; and
- spatial strategies (such as varietal mosaics and mixtures) have an important role to play.
The situation that causes particular concern is highlighted by Blackleg in canola. In this case, there are a number of resistance genes in circulation in different canola varieties. However, with just one exception they have all broken down at least in some areas.
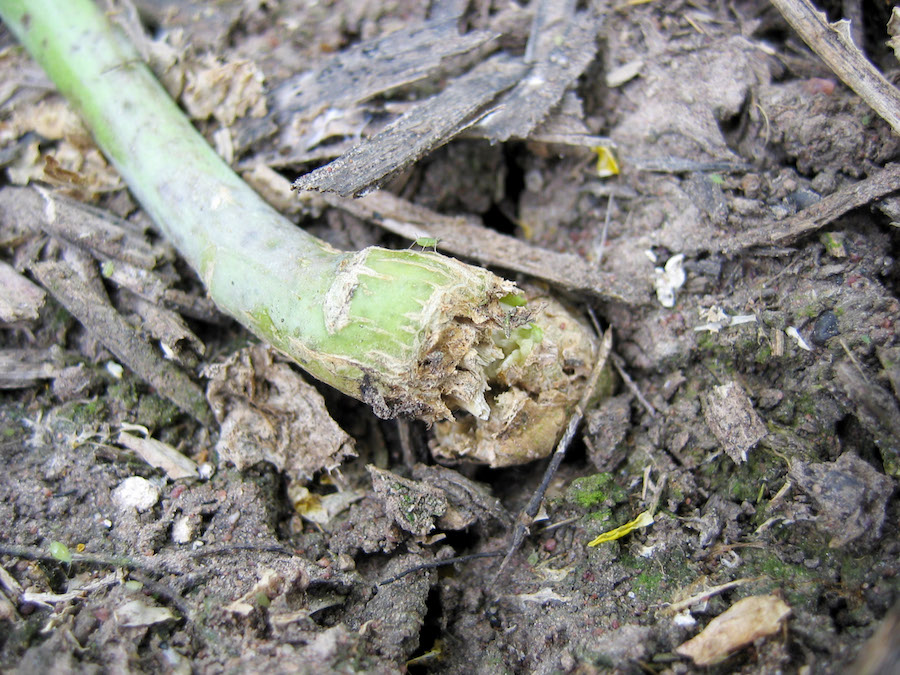

Blackleg (Leptosphaeria maculans) in canola. Photo: S Marcroft
As Dr Barrett says: “Many canola varieties still carry four or five of these resistance genes. From an evolutionary perspective, this is almost the worst thing you can do, to continue to cultivate these genes, because it basically encourages the evolution of super-pathogens that can overcome all resistance genes available.”
However, modelling and experimental work show there is potential to extend the use of resistance genes. This becomes especially important given that the world does not have an inexhaustible supply of novel resistances.
“Finding a new resistance trait takes a lot of time, effort and money,” Dr Thrall says. “So, this is a resource we want to conserve as much as possible. That means using resistance genes in a way that maximises their effectiveness and longevity.”
Solutions
Using resistance genes as a renewable resource brings to light two major considerations:
- the importance of spatial or temporal rotations of these genes – referred to as ‘strategic rotations’; and
- strategic rotations requiring landscape-scale planning that exceeds the farm scale.
“Being very clever with rotations and awareness of the gene/pathogen mixes in the landscape is the way forward,” Dr Thrall says.
This amounts to a more complicated version of the basic rotation strategy currently in use. It involves not just changing the resistance type in time but moving where the resistance is in the landscape in a coordinated fashion.
“Rotations are a strategy that seem to come up as a pretty good approach,” Dr Thrall says. “It’s about presenting a pathogen with a continually changing mix of resistance in space and time to keep the pathogen off balance, but in a way that is also feasible for a grower to implement.”
A key challenge is the level of collective coordination across many farms that is needed to reap the benefits. That brings with it a new kind of challenge. As Dr Thrall says: “The one example of this being done successfully is Bt (Bacillus thuringiensis) resistance in cotton, where grower were legislated to protect the resistance gene by planting refuge crops to minimise evolution in the insect pest. There’s currently nothing that works like that for fungal pathogen and crop interactions. But that’s the direction we need to go.”
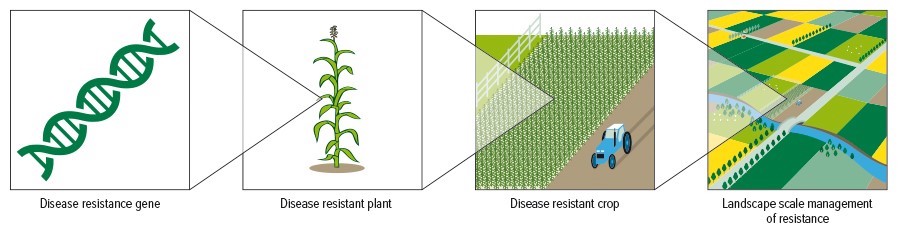
Strategic rotations require coordinated deployment of disease resistance genes at the landscape scale. Source: CSIRO Agrilculture & Food
Having more crops available that are profitable to include in a rotation would ultimately also serve to disrupt the evolutionary process or reduce the pathogen population size.
Achieving this more ambitious level of disease control innovation will ultimately become imperative given a hard fact about host-pathogen evolution. As Dr Barrett puts it: “If there is a resistance gene out there, it is very likely that somewhere in the world there is a pathogen that can overcome it.”
Paddock realities
While the simulations point the way to the ideal disease control scenarios, the CSIRO researchers are well aware that work is needed to make these realistic for the grains industry.
In response, they are now working a new capability into the modelling to deal with practical economic factors that affect farm profitability, such as input costs. They also want to build in the impact of management interventions such as fungicide use.
This is part of broader efforts looking at barriers to coordinated action within landscape-scale strategies that includes:
- implementation challenges faced by growers;
- ensuring the varieties needed for a successful strategic rotation are available;
- ensuring breeders have access to information they need to breed varieties compatible with strategic rotations; and
- strategic monitoring of the pathogen population.
“Strategic rotations at a landscape scale are achievable but there are hurdles to overcome,” Dr Thrall says.
Surveys of what breeders, agronomists and growers would be willing to do differently to try to maximise the life of resistant varieties got underway in 2020. Until that information is integrated, the researchers have a key message: the way we deploy genes in landscapes matters.
Benefiting from evolutionary insights, however, will require people to think in a broader and more cooperative way about the deployment of resistance genes. That includes the willingness to pull back varieties for the greater good.
“Ultimately, it is about thinking more cooperatively to get more value for everyone from these resources,” Dr Thrall says.
Dr Barrett agrees. As with antibiotic and herbicide resistance, these are predictable events yet these valuable resources were deployed without building in protection to ensure their longevity. “There’s a step change needed where we are more proactive from the outset by including an evolutionary perspective about how we develop, deploy, use and withdraw resistant cultivars,” he says. “Discovering more disease resistance genes just gives us more pieces to move around on the chessboard. How we use them remains the frontline.”
He is also keen to address the perception of modelling as abstract and distanced from the real world.
“We are not ivory tower thinkers,” Dr Barrett says. “Pete grew up on a dairy farm in Connecticut in the US and I worked on the family farm in the Western Australian wheatbelt. So, we understand the reality of farming and the people we work for.”
More information: Peter Thrall, peter.thrall@csiro.au; Luke Barrett, luke.barrett@csiro.au.

























































